Why There’s Rarely “A Gene For” Anything
Genetics coverage keeps finding 'a gene for' everything. The science tells a more complicated story. A closer look at what that framing gets wrong—and why it matters.
Why "a gene for" language dominates science headlines
Spend any time reading science coverage and the pattern becomes familiar. Researchers have identified a gene for obesity. A gene linked to alcoholism. A gene for aggression, for loneliness, for how well you sleep. The phrasing is simple and intuitive. It suggests that complex human traits can be traced to a single biological switch—a specific piece of DNA that explains why we are the way we are.
It’s almost always misleading.
This language has been a consistent feature of genetic reporting since at least the 1990s, when the Human Genome Project generated enormous public interest in what DNA might reveal about human nature. The field of genetics was advancing rapidly, and early discoveries in single-gene conditions suggested that hereditary diseases might yield straightforward biological conclusions.
The “a gene for” framing persisted because it works rhetorically. It implies that researchers found the thing—the specific cause of a specific outcome. It maps onto how people already think about inheritance: you got your father’s nose, your grandmother’s temperament, your family’s predisposition to heart disease. Genes feel like explanations.
The problem is that for most traits, this model is wrong. Not just partially wrong or overstated, but structurally wrong. The biology doesn’t work that way, and understanding why changes how much weight any individual genetic finding deserves.
What genes actually do
A gene is a segment of DNA that contains instructions for making a protein. Proteins perform much of the functional work in cells: they form structures, transmit signals, catalyze reactions, and regulate other genes. The connection between a gene and an observable trait runs through this chain, and it’s rarely short or direct.
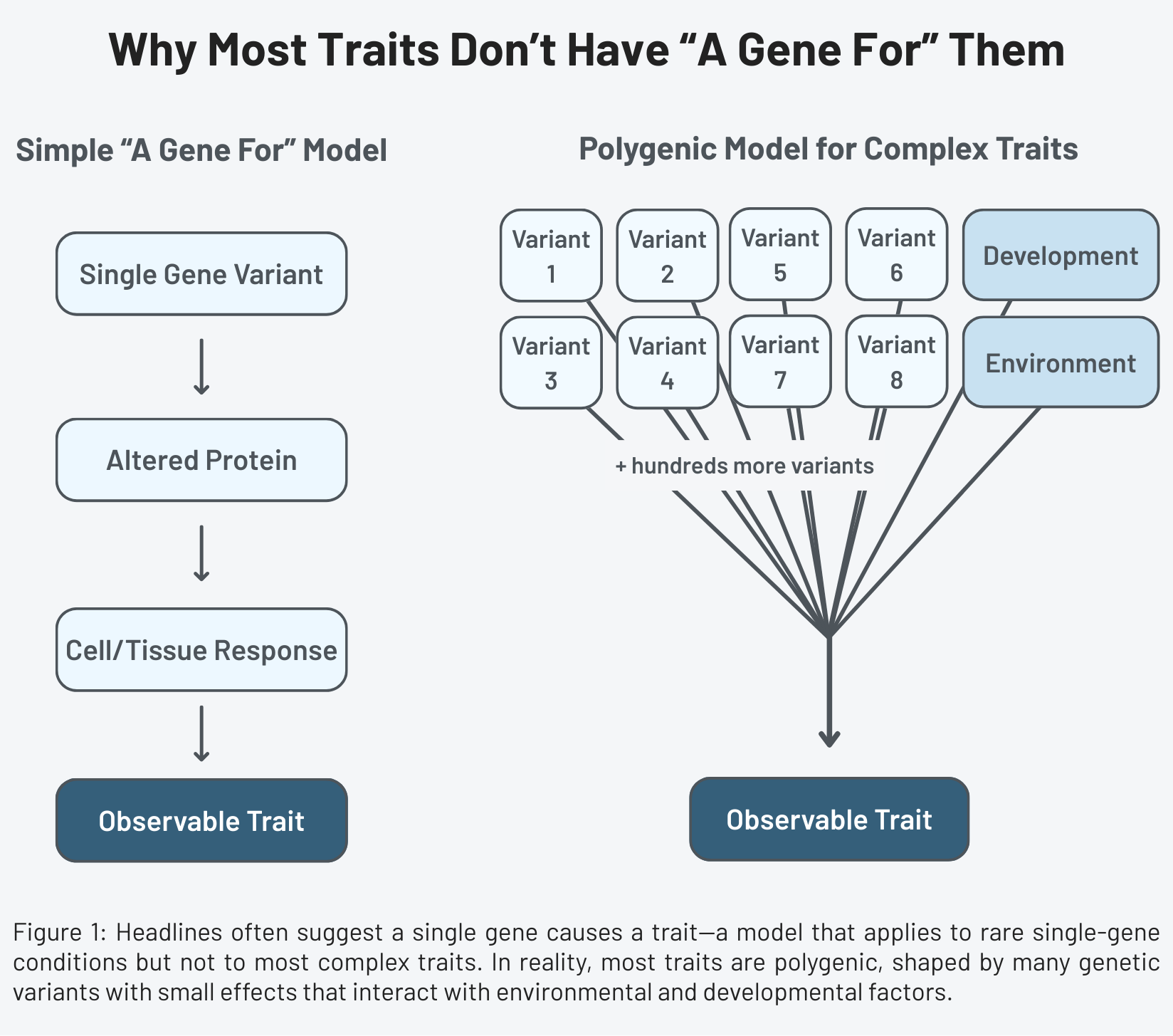
Consider height. Height is heritable: tall parents tend to have taller children. But height isn’t determined by one gene producing one outcome. It reflects the combined effects of thousands of genetic variants interacting with one another and with environmental factors, each influencing processes like hormone signaling, nutrient absorption, and bone growth. A single variant associated with height might shift average height by only a few millimeters across a population.
This is what geneticists call a polygenic trait: a characteristic influenced by many genes. The vast majority of human traits are polygenic. Some involve thousands of variants spread across the genome. The idea that any one of these variants constitutes “the gene for” the trait misrepresents the underlying biological architecture.
Carrying a particular genetic variant associated with a trait is not the same as having a deterministic cause for that trait. At most, it represents one contributing factor.
The difference between single-gene and complex-trait genetics
Some conditions are monogenic. Huntington’s disease is caused by a specific mutation in a single gene. If an individual carries a pathogenic expansion above a certain threshold, they will develop the disease. The relationship is direct and highly predictable. Cystic fibrosis, sickle cell disease, and a number of other rare disorders follow similar logic: a single pathogenic variant plays a central causal role.
These disorders represent a small subset of human diseases, and they’re precisely the cases that gave early genetics its cause-and-effect reputation. That reputation then extended into research on complex traits, where it does not apply.
Intelligence, depression, type 2 diabetes, schizophrenia, and longevity do not work like Huntington’s disease. Genome-wide association studies (GWAS), which scan hundreds of thousands of genetic markers across large populations, have identified dozens to thousands of variants associated with most common diseases and complex traits. Not one variant—thousands.
For major depression alone, large-scale studies have identified more than a hundred genetic variants with statistically significant associations to risk, and each one explains only a tiny fraction of the variance. The same pattern holds for educational attainment, body mass index, coronary artery disease, and nearly every behavioral or psychiatric trait studied at scale.
A headline about “the gene for depression” implies a biological on/off switch. The research describes modest shifts in probability distributed across the genome, mediated by development and environment, that cannot be reduced to a single variant without distorting what the science shows.
Gene expression isn't fixed
Even when someone carries a variant associated with a particular trait or condition, whether and how that variant influences outcomes depends heavily on context—something the “a gene for” framing ignores.
Genes don’t function in isolation. They're regulated by other genes (epistasis), by cellular signaling pathways, by developmental timing, and by chemical modifications to DNA and its associated proteins—processes referred to collectively as epigenetic regulation. These modifications influence whether a gene is expressed and how much protein it produces, without altering the underlying DNA sequence. They can be shaped by stress, diet, environmental exposures, and age.
This means the same DNA sequence can function differently in different people, in different environments, and at different stages of life. A variant associated with elevated risk under one set of circumstances may have little or no measurable effect under another set of circumstances. Gene-environment interactions are central to how genetic variants relate to traits.
Identical twins illustrate this clearly. They share nearly identical genomes, yet they often differ in whether they develop conditions with strong genetic components, including schizophrenia, type 1 diabetes, and bipolar disorder. If genes determined outcomes directly, this wouldn’t occur. The fact that it does underscores how biological context shapes what any given DNA sequence ultimately produces.
How association gets mistaken for causation in genetics reporting
Genome-wide association studies identify statistical relationships between genetic variants and traits across large populations. A particular variant may appear more frequently in individuals with a given condition than in those without it. That association is meaningful because it points researchers toward biological pathways worth investigating, but it’s not the same as establishing that the variant directly causes the trait.
Many associated variants are located in non-coding regions of the genome. They don’t change the structure of a protein. Instead, they may influence when, where, or how much of a protein is produced—or they may simply be located near the true causal variant. Identifying an association is the beginning of mechanistic investigation, not the conclusion.
The “a gene for” framing collapses this distinction. It takes a finding that says “this variant is statistically more common in people with this trait” and translates it into “this variant causes this trait in this individual.” These are fundamentally different claims, and the evidence rarely supports the latter.
Population-level statistics also don’t translate into individual predictions the way the framing implies. A variant associated with increased risk—even substantially increased risk—typically shifts probability rather than determining outcome. Most people who carry a given risk variant will never develop the associated condition. Many people who develop the condition will not carry the specific variant highlighted in the headline.
If you want a faster way to read health studies without oversimplifying them, subscribers get a free guide: How to Read a Health Study in 10 Minutes.
When the framing does fit (and when it doesn't)
The “a gene for” language is not always inappropriate. In certain contexts, it is accurate and clinically meaningful.
Monogenic disorders are the clearest example. Pathogenic variants in BRCA1 or BRCA2 substantially increase the lifetime risk of breast and ovarian cancer. Hereditary cancer syndromes, certain rare metabolic disorders, and specific pharmacogenetic variants that influence how individuals metabolize medications—in these cases, identifying a particular genetic variant carries significant predictive value and can directly inform clinical decisions.
The problem arises when this logic is applied to complex behavioral or psychological traits, where the underlying biology is fundamentally different. A BRCA1 variant meaningfully alters cancer risk in a way that can guide screening and preventive strategies. A variant associated with introversion in a genome-wide association study adds almost nothing to predicting whether a particular person will be introverted.
A frequently cited example of this misapplication is the MAOA gene, sometimes labeled the “warrior gene” in popular coverage. Certain variants of MAOA have been associated in some studies with increased aggressive behavior, particularly in individuals who experienced childhood adversity. Media coverage often implied a direct causal link between carrying the variant and being predisposed to violence.
The research is more limited and more conditional than that framing suggests. The findings describe statistical associations observed in specific populations under specific environmental circumstances, with ongoing debate about the effect sizes and the mechanisms involved. The variant doesn’t program aggression. Under certain developmental conditions, it may influence neurochemical pathways in ways that affect behavior at the population level. That is a much narrower and more accurate claim than what most headlines conveyed.
How to read genetics headlines more carefully
When you encounter a headline announcing the discovery of “a gene for” something, a few questions can help clarify what the finding actually shows:
- Is this a monogenic trait or a complex one?
- Does the study report an association, or has a causal mechanism been established?
- How large is the effect size? Does it meaningfully shift predictions at the individual level?
- Has the finding been replicated in independent, large-scale samples?
A headline that reads “Scientists find the gene for stress resilience” can be more accurately framed as: “Researchers identify genetic variants associated with stress response, each with small effects and influenced by environmental context.” The latter is less dramatic. It’s also closer to what the evidence usually supports.
Why this matters
Misunderstanding genetic determinism has consequences beyond science literacy.
Genetic fatalism is one risk. People who believe they carry “the gene for” a disease may conclude that prevention is pointless—that the outcome is biologically inevitable regardless of behavior. For most complex conditions, that conclusion is not supported by the evidence. Lifestyle, environment, and chance interact with genetic predisposition in ways that preserve meaningful opportunity for intervention.
Stigma is another concern. Attaching traits such as aggression, addiction, or intelligence to specific genes—in the simplified way popular coverage often does—implies that these characteristics are fixed, biologically determined, and located entirely within the individual. That framing ignores the role of social and environmental context, and it has a history of being misused. The underlying science doesn’t justify it.
At the consumer level, direct-to-consumer genetic testing can reinforce these misunderstandings. Some services present trait-associated variants as personal risk factors without adequately conveying effect sizes, population-level uncertainty, or the substantial limits of individual prediction.
None of this diminishes the importance of genetic research. Genome-wide association studies have meaningfully advanced understanding of disease biology. Identifying genomic regions associated with conditions helps researchers prioritize pathways for investigation. Polygenic risk scores, when carefully validated and appropriately contextualized, are beginning to show utility in specific clinical settings.
But the science is more complex than the headlines suggest. When a modest statistical association is presented as a discrete biological cause, it shapes what patients expect, what they fear, and what decisions they make in ways that outpace the evidence.
The more accurate picture is less tidy. Most human traits reflect the interaction of many genetic variants, environmental influences, developmental processes, and chance. They do not operate through a single switch.
Recognizing that distinction doesn’t make genetics less powerful. If anything, it makes it more so. The polygenic architecture of complex traits, the interplay between genes and environment, and the dynamic regulation of gene expression reveal a biological system that is intricate, context-dependent, and still being understood.
That complexity is harder to compress into a headline. It is also closer to what the research actually shows.
References
- The gene: An appraisal, Progress in Biophysics and Molecular Biology
- The polygenic and poly-environmental nature of personality, Current Opinion in Psychology
- Human Height: A Model Common Complex Trait, Annals of Human Biology
- Twenty years of genome-wide association studies, European Journal of Human Genetics
- Genome-wide association studies: the good, the bad and the ugly, Clinical Medicine
- Huntington disease: Advances in the understanding of its mechanisms, Clinical Parkinsonism & Related Disorders
- BRCA1- and BRCA2-Associated Hereditary Breast and Ovarian Cancer, GeneReviews
- The role of monoamine oxidase A in aggression: current translational developments and future challenges, Progress in Neuro-Psychopharmacology & Biological Psychiatry

